Phylogenetics
Phylogenetics is the science of the evolutionary relationships among species. It is best visualized in tree-strucutre where the branch lengths correspond to some notion of evolutionary time.
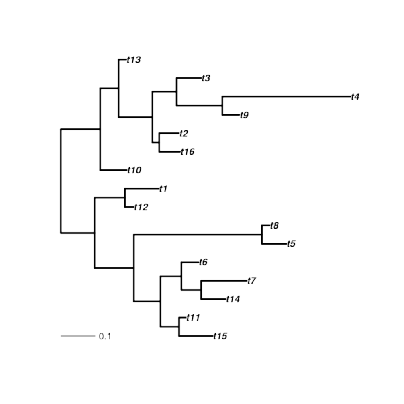
Assuming a trait is measured across the tips/leaves of a phylogeny, how can one infer estimates about the values of trait at some ancestral states?
Given a number of traits, observed across the members of a phylogeny, how can you distinguish which traits are due to phylogenetic evolution and which are not (eg. are due to enviormental adaptation)?
