Meera Unnikrishnan
Technical Summary
Our group is interested in understanding how bacterial pathogens interact with the host using a combination of whole genome-based strategies and single gene-level functional studies.
Bacteria have developed several strategies to survive within the host, including intracellular survival and formation of biofilm communities, enabling them to escape antibiotics and immune responses. Investigating such mechanisms, both from the bacterial and host standpoints, is crucial for the development of effective preventive and therapeutic approaches, particularly for recurrent bacterial infections. Our key research focus is on understanding host-bacterial interactions that are key to persistent infections. Our recent research has also centered on understanding the role the microbiota plays in modulating host-pathogen interactions.
Major areas of interest in the group include:
Colonisation of the gut by Clostridium difficile:Interactions of nosocomial pathogen C. difficile with the gut epithelium and with the gut microbiota are poorly understood. The roles of many extracellular and surface proteins of C. difficile in colonisation and establishment of infection remain unclear. We have previously identified bacterial factors, including secreted proteins that may play important roles during gut infection. Additionally, we have characterized processes such as C. difficile biofilm formation and proteins that modulate bacterial community formation in vitro. Current work in the group focuses on understanding bacterial mechanisms underlying C. difficile colonisation and persistence. One of our current interests include development of new cell-based systems to enable pathogen-host-microbiota interactions.
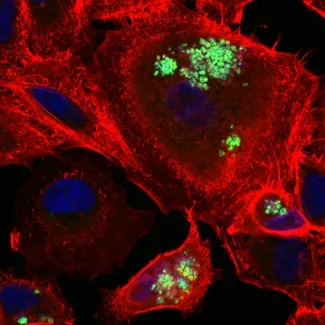
Intracellular survival of bacterial pathogens: Intracellular pathogens are hard to target, particularly in bone infections. Diagnosis of bone infections is also problematic, with new markers urgently needed. Current work focuses on understanding bacterial factors important for intracellular survival of S. aureus and Mycobacterium tuberculosis and developing antimicrobials that target intracellular pathogens. In collaboration with the Perrier group in Chemistry we are developing nanoparticle-based approaches to delivering antibiotics to effectively kill intracellular bacteria.
Type VII (Esx) secretion systems of Staphylococcus aureus: The intriguing staphylococcal type VII secretion systems secrete virulence factors that have excellent vaccine and drug potential. However, their functions are during infection are not clear. Our previous work has shown a role for the staphylococcal Esx proteins in modulating intracellular infection. We are currently examining how these specialized systems mediate interactions with host cells to modulate immune responses and their roles in bacterial physiology.
Selected publications:
1. Tchoupa, A.K., Watkins K.E., Jones, R.A., Kuroki, A., Alam, M. T., Perrier, S., Chen, Y., Unnikrishnan, M. (2019) The type VII secretion system protects Staphylococcus aureus against antimicrobial host fatty acids. Scientific Reports Sci Rep 10,14838 (2020).
2. Anonye, B.A., Hassall, J., Patient, J., Detamornrat, U., Aladdad, A.M., Schüller, S., Rose, F.R.A.J., Unnikrishnan, M. (2019) Probing Clostridium difficile infection in complex human gut cellular models. Frontiers in microbiology 10, 879
3. Korea, C.G., Balsamo, G., Pezzicoli, A., Merakou, C., Tavarini, S., Bagnoli, F., Serruto, D.,and Unnikrishnan, M. (2014) Staphylococcal Esx proteins modulate apoptosis and release of intracellular Staphylococcus aureus during Infection in epithelial cells. Infect. Immun. 82(10): 4144-53
4. Dapa, T., Leuzzi, R., Ng, Y.K., Baban, S., Kuehne, S., Scarselli, M., Minton, N.P., Serruto,D., Unnikrishnan, M. (2013) Multiple factors modulate biofilm formation by the anaerobic pathogen Clostridium difficile. J. Bacteriol. 195(3): 545-55

