Software documentation and notes
Software development
The initial phase of the work involved significant amounts of work on the software required within the System Biology group to process and analyse genomic data.
The software makes considerable use of open source software and libraries, much of which is excellent, but often poorly documented. Much of what I learnt was from individual websites and blogs where users had posted additional helpful tips and examples. The following is my attempt to add to this fund of knowledge.
Installing and building software on Windows PCs:
Installing Perl on a Windows PC
This is relatively easy, although there are one or two things to watch for, so some notes on installing Perl on a windows PC
Perl is used significantly within the BioInformatics community, which uses the BioPerl library and the Perl based API for accesssing Ensemble genetics data
Installing Visual C++ Express, and creating mixed .NET and classic C++ applications
First, some notes on installing Visual C++ express.
The recent, free, versions of Visual C++ are heavily oriented towards using Microsofts .NET architecture with data held in managed memory. There are many situations where a development needs a mixture of .NET and 'conventional' C++, such as when there is a need to embed a large amount of existing code into a new application.
While Microsoft make it easy to develop a pure .NET application, a mixed .NET and classic C++ application is more difficult, so I have made some notes which may be of help to others faced with the same problem.
This also includes some notes on problems with the OpenFileDialogBox freezing when the dll is used on Vista.
Installing OpenSSL on Windows
Instruction on how to to install OpenSSL from the source, and build a static library, which turns out to be very easy
Installing and building software on Apple MACs
Problems building code that uses the Carbon library
Some open source software makes use of the Carbon library, and typically provides no information as to how to configure to link to the appropriate libraries. This is what needs to be done.
Installing gcc and boost on an Apple XServe platform
The darwin OS that Apple provides for its XServe platform includes a C/C++ compiler that is based on gcc 4.0.1. This is over 4 years old and I have documented how I installed a local, later release of the gcc compiler and then installed boost to work with this compiler rather than the Apple compiler.
The reason for installing these updates was that prior to release 4.2, gcc is unable to cope with anonymous namespaces in precompiled headers, and boost makes extensive use of anonymous namespaces in its libraries, so the compiler that comes on the Apple platform prevents their inclusion in precompiled headers, which is something of a limitation.
Notes on using Perl
Perl, its efficiency and architectures involving mixed Perl and compiled code.
While Perl has many, well documented advantages, I was concerned that, as a scripting language it was likely to be very inefficient, indeed the BioPerl website notes that people frequently comment that BioPerl is slow.
I decided to compare the efficiency of some library code from the BioPerl website with the same algorithm implemented in C++. Three different tests were performed, and C++ was 150, 10 and 1.1 times faster.
I felt that what would be ideal would be an architecture that allowed a mixture of compiled code (for efficiency) together with Perl code (for ease of development)
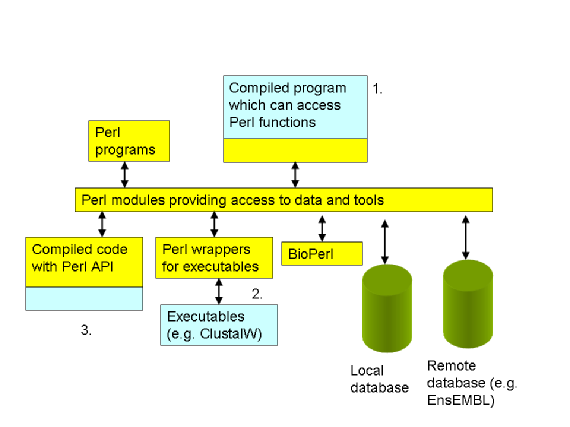
There are at least three ways in which Perl and compiled code can be used together:
-
A program is written in a language such as C or C++ which can use Perl when required, giving it access to other systems that have a Perl API. I have created an example of such a program, which has been tested on Windows and MacOSX to demonstrate how this can be done.
-
A Perl module can be written which runs an executable and returns the results. This is relatively simple and BioPerl contains many such examples, eg modules for running ClustalW.
-
A Perl module can be created where core functions are provided by a compiled library (a dll in Windows, a dylib file on OSX) that is embedded within the module. This is how many of the standard modules within Perl have been created, e.g. those that provide database access. It is perhaps significant that no BioPerl modules have been created in this way, suggesting that they may be inappropriate for performing operations that involve significant amounts of data.
This is covered in more detail in the following section
Embedding C code into Perl
C code that has been embedded into Perl is called an XSUB. Perl provides a set of scripts that will create a directory with template files that can be extended to provide the functionaility required, and which can then be built, also using Perl scripts which make and use the appropriate makefiles.
This is covered, with examples, in the Perl documentation
It is also covered in other web sites such as O'Reilly's "Programming Perl'
An alternative approach is to use SWIG which automates, as far as possible, the process of wrapping C/C++ code such that it can be used by other languages such as Perl.
