Experimental Biology Project
Control of Reactivity of Protein-Disulphide Isomerase.
Dr Katrine Wallis, Life Sciences (Formerly Biological Sciences).
Structural Biology - Robert Freedman Group
Project Background:
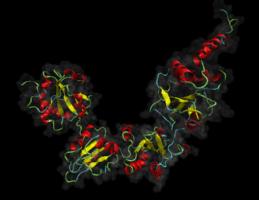
Proteins exported from the eukaryotic cell undergo a complicated folding and quality control pathway before they are ready to leave the endoplasmic reticulum. An important part of this folding pathway is the introduction of disulphide into the proteins; without these many exported proteins are unable to reach their native state. The formation of correct disulphides is a slow process but it is catalysed by enzymes belonging to the protein-disulphide isomerase family. The most well-known of these is protein-disulphide isomerase (PDI) itself but the family consists of at least 19 proteins. These proteins differ in their expression pattern and architecture as well as their reactivity towards disulphides and substrate specificity. Generally their exact biological function is not understood and the molecular details determining their reactivity and substrate specificity not well characterised. The aim of this project is to understand the difference between two of these proteins namely PDI and its close homologue PDIp (pancreatic PDI), which is only half as active as PDI in standard assays. The two proteins share the same domain architecture abb’xa’c where the a and a’ contain an active site each and the b’ domain contains the main substrate binding site. Hence it is possible to determine effects on reactivity arising from active sites differences and substrate specificity independently of each other.
We aimed to test two hypotheses for the low reactivity of PDIp:
- The difference in reactivity is due to one of the two active sites in PDIp being CTHC rather than CGHC as found in both PDI active sites and the first PDIp active site, with the larger threonine residue causing steric hindrance for the formation of a disulphide between the two cysteines needed for the activity. This will be tested by making the T->G mutant in the non standard active site of PDIp.
- The lower reactivity of PDIp is caused by differences in the substrate recognition stemming from the low identity between the two b’ domains (38% in b compared to 50% identity in the full length protein). This can be tested by swapping the substrate binding domain in PDIp for the substrate binding domain in PDI. This swapping will be done by primer extension pcr in two steps, first creating the chimera abb’xa’c (where PDIp domains are in black and PDI domain in red) and then using this chimera as a template to create abb’xa’c. Once the new expression constructs are made the proteins need to be expressed in E.coli and purified. We already have established protocols for expression and purification of both PDI and PDIp involving immobilised metal affinity and ion exchange chromatography and are expecting the new constructs to behave similarly to the wild type enzymes. We will then test reactivity of these new proteins using standard assays for PDI activity such as reduction of disulphides in insulin and the ability to refold a small disulphide rich protein substrate, RNase.
Further Reading:
Hatahet, F. & Ruddock, L.W. (2007). Substrate recognition by protein disulphide isomerases. FEBS. 274, 5223-5234.
Conn, K.J., Gao, W., McKee, A., Lan, M.S., Ullman, D., Eisenhauer, P.B., Fine, R.E. & Wells, J.M. (2004). Identification of the protein disulphide isomerase family member PDIp in experimental Parkinson's disease & Lewy body pathology. Brain Research, 1022, 164-172.
(*Project Background adapted from the original MOAC mini-project proposal)