PhD topic: DNA-templated synthesis
The initial brief of my PhD project decided October 2011.
The PhD project will involve using DNA-templated assembly to investigate controlled manufacture of polymers and other molecules, and their properties. The study will begin by looking at building a conjugated polymer and to investigate any advantageous electrical properties. The challenge will be to build the monomers in the polymer in a ‘trans’ formation relative to each other. It is not known if the current methods already used by the groups in Warwick and Oxford will allow repeated trans bonding to occur, and a new approach may need to be developed be to join the DNA-attached monomers in the desired configuration.
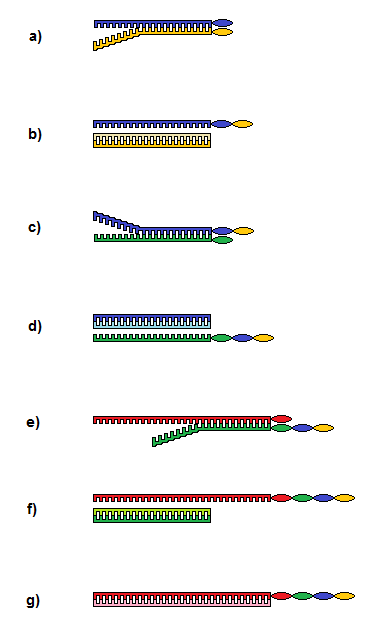 Figure 1:
Figure 1:
a) Strand 1 (S1-yellow) and strand 2 (S2-blue) come together and hybridization occurs along the majority of the strands, but leave a ‘toehold’ section un-hybridized: this will enable separation using complementary strands.
b) Group transfer has happened at the strand ends from S1 to S2, and S1 is removed using hybridization with a complementary strand C1.
c) Strand 3 (S3-green) now hybridizes with S2 ready for the end groups to be transferred.
d) S2 is now removed using complementary strand C2, leaving a string of 3 groups now on S3.
e) Strand 4 (S4-red) now repeats the process used so far. S4 is the last strand in the sequence and is longer than the previous strands.
f) S3 is removed with C3 and the designed string of groups are left on the end of S4.
g) A complementary strand C4 is hybridized with S4. This will enable easy detection of the strand using PAGE.
Steps b) to d) can be repeated as needed to create a string with a desired sequence of monomers.
My two supervisors for the project are:
I am also grateful for the support of my advisory committee:
Dr Corrine Smith
