MSc Project 1
Novel Biophysical Methods for Membrane Protein Research
Supervised by Dr. David I. Roper
Background
The biosynthesis of bacterial cell wall peptidoglycan involves a complex pathway of cytoplasmic, membrane-associated and extracellular stages. The intercellular stages culminate with the production of the membrane-anchored precursor, Lipid II.1 Newly synthesised Lipid II is then translocated across the inner membrane to the extracellular side by a currently unknown mechanism. The translocation process has recently been identified as a property of the protein FtsW with indirect involvement from the inner membrane protein MurJ (MviN), both of unknown structure.2,3
In this study, FtsW and MurJ were inserted into Green Fluorescent protein (GFP)-fusion vectors which are ideal systems for expression, isolation and biophysical studies of proteins. Isolation of membrane proteins for structural and functional studies is a notoriously difficult procedure often generating only low yields of expressed protein. In addition, membrane proteins usually exist within the context of a lipid bilayer and at present require extraction by detergents providing an artificial lipid environment that may disrupt its native structure.
Styrene maleic acid lipid particles (SMALPs) are nanoscale “cookie cutters” that have the potential to isolate a protein within its natural lipid environment (Fig. 1).4 When amphiphatic particles, such as the synthetic copolymer Styrene Maleic Acid (SMA), interact with lipids they assemble into nanoscale discs. A membrane protein can be wrapped up within the lipid disc causing it to be isolated whilst remaining in its native lipid environment. This simple method preserves the native structure and activity of the protein and eliminates the need for detergent solubilisation. Initial experiments using the GFP expression system have suggested that the SMALPs lipid disk shields the N or C-terminus polyhistidine tag thus preventing purification by immobilised metal ion affinity chromatography (IMAC). We exploited a new multifunctional GFP (mfGFP) in the fusion vector in which the histidine tag exists on a loop within GFP.5 We proceeded to generate expression of the proteins FtsW and MurJ with GFP or mfGFP fluorescent tags so they would be readily available for isolation and purification by SMALP extraction and IMAC purification. Results from this mini project will provide a starting point towards developing a comprehensive understanding of the Lipid II translocation mechanism and potentially exposing FtsW and MurJ as new targets for antibiotics.
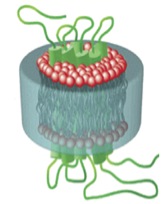
Fig. 1: 3D Model of a protein and lipids wrapped up inside a SMA particle produced by SMALP solubilisation.
References
[1] Barreteau H, Kova A, Boniface A, Sova M, Gobec S & Blanot D (2008) Cytoplasmic steps of peptidoglycan biosynthesis. Fems Microbiology Reviews 32: 168-207.
[2] Mohammadi T, van Dam V, Sijbrandi R, et al. (2011) Identification of FtsW as a transporter of lipid-linked cell wall precursors across the membrane. The EMBO Journal.
[3] Ruiz N (2008) Bioinformatics identification of MurJ (MviN) as the peptidoglycan lipid II flippase in Escherichia coli. Proceedings of the National Academy of Sciences of the United States of America 105: 15553-15557
[4] Knowles TJ, Finka R, Smith C, Lin YP, Dafforn T & Overduin M (2009) Membrane proteins solubilized intact in lipid containing nanoparticles bounded by styrene maleic acid copolymer. Journal of the American Chemical Society 131: 7484-7485.
[5] Kobayashi T, Morone N, Kashiyama T, Oyamada H, Kurebayashi N & Murayama T (2008) Engineering a novel multifunctional green fluorescent protein tag for a wide variety of protein research. PloS one 3: e3822.
