Mini-Project 1
Molecular Dynamics Investigation of Biomimetic Antifreeze Macromolecules
Supervised by Dr. Rebecca Notman
Antifreeze glycoproteins are essential for preventing ice-induced death in polar fish. They function at much lower concentrations than conventional antifreeze agents, such as DMSO and glycerol. These proteins could improve cryostorage of biological materials, such as blood, stem cells and potentially whole organs. These proteins are not exploited due to the paucity of the material, this is a result of difficulties in the extraction of significant quantities from natural sources and the challanges of the total synthesis. Recently, the Gibson Group have demonstrated that relatively simple synthetic polymers can reproduce some of the properties of these proteins. These are 3 peptides functionalised with a monosaccharide. The aim of this project is to use molecular dynamics (MD) to elucidate the molecular-level mechanisms of these polymers.
All atom simulations of 4 molecules were carried out for 100 ns in a specially selected 4 site explicit water model, TIP4P/ICE. This model was chosen as it gives the best representation of the melting piont of water. The simulations were run at two temperatures above and below the freezing of water. The two temperatures were 267 K and 298 K which correspond to the temperatures used in the Gibson lab for recrystallisation inhibition testing.
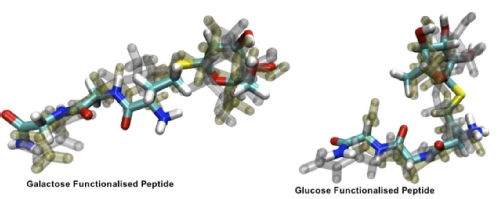
The Gibson groups found that the tripeptide functionlised with galactose showed appreciable recrystallisation inhibition, whereas the tripeptide functionlised with the stereoisomer of galactose, glucose did not. The most dominant structures determined by cluster analysis from the MD simulations showed a marked differenece in the structures at 267 K. The pictures show the three most dominant structures of each of the molecules overlaid. The galactose functionalised peptide showed a more extended solvent accessible structure.
Here is the Presentation I gave to the Notman group about my mini-project.
